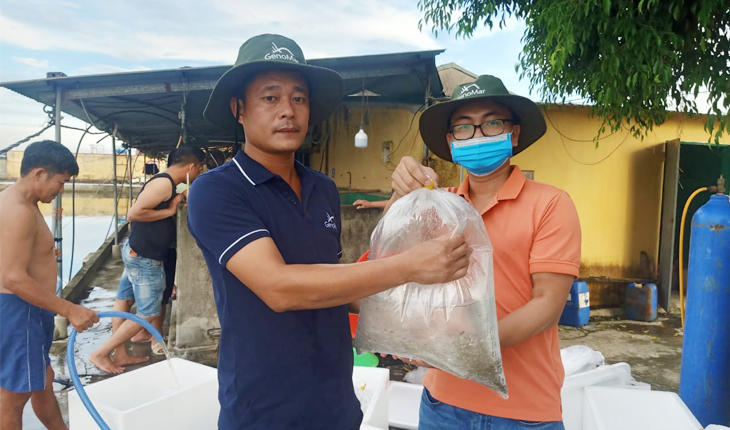
Development of new real-time PCR methods for detection of Decapod iridescent virus 1 in shrimp
Genics’ R&D Team has produced new research on Decapod Iridescent Virus (DIV1) real-time PCR assays published as free open access for all in the global shrimp industry. This scientific research article is a timely reminder for continuous shrimp pathogen surveillance and disease mitigation via proven next-generation technologies such as Genics’ Shrimp MultiPathXtra.
Decapod Iridescent virus (DIV1) infections emerged in mainland China around 2014 and have devastated shrimp aquaculture operations in Chinese coastal provinces. In 2020, DIV1 has spread to Taiwan with devastating results to shrimp and crayfish farms, in addition to being found in wild caught Penaeus monodon from the Indian Ocean. This trend is a major cause for concern and an urgent reminder to expand the tools needed to monitor the spread of DIV1 globally.
Trials
A set of four different real-time polymerase chain reaction (PCR) assayswere positioned across the genome of DIV1 to detect the virus in shrimp tissues. All four assays show a wide dynamic range and high analytical sensitivity and specificity. In addition, the newly developed assays show excellent diagnostic sensitivity and specificity in clinical Litopenaeusvannamei samples of North Asian origin. The new molecular toolset will enhance global capabilities to monitor the spread of DIV1 and ultimately be used as an early warning system for farmers and authorities to engage in appropriate risk mitigation strategies.
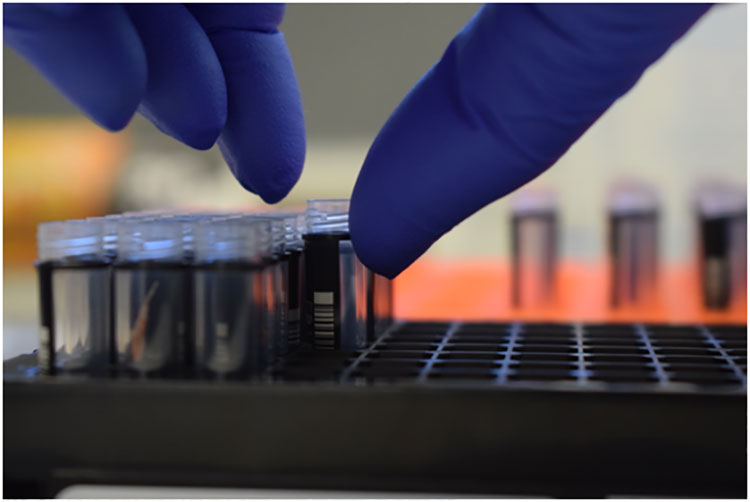

Vannameipleopod samples were collected from live individual shrimp taken from an earthen pond suffering mortalities in Northern Asia. Shrimp samples were submerged in 70% Laboratory Grade Ethanol for preservation and sub- sequent total nucleic acid (TNA = RNA and DNA) extraction. TNA was extracted using a MagMAX™ Core nucleic acid purification kit with the KingFisher FLEX robot (Thermo Fisher Scientific, CA) as described by Moser, Franz, Firestone, and Sellars (2022). Extracted sample TNA was eluted in 50 μl PCR grade water and directly used for the different analyses.
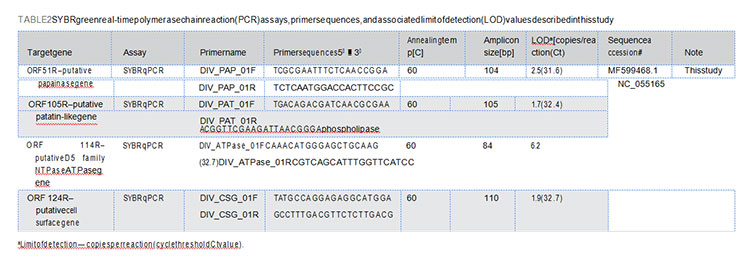
Real-time PCR assays were setup in 384-well PCR plates using the PowerUp SYBR Green Master Mix and run on the QuantStudio 12K Flex Real-Time PCR system (Thermo Fisher Scientific, CA) described elsewhere (Moser et al., 2022).
Real-time PCR amplicons were sequenced directly to confirm sequence authenticity. PCR amplicons in original ampli- fication reaction mix/volume and corresponding forward and reverse primers (3.2 μM) were submitted to the Australian Genome Research Facility (AGRF, Brisbane Australia) for direct Sanger sequencing using Big Dye Termina- tor chemistry 3.1 and ABI Capillary Sequencer 3730xl (Thermo Fisher Scientific, CA).The quality of sequence traces was checked and curated using the Sequencher Software (GeneCodes, MI) and verified sequences confirmed using the BLAST tool interface of NCBI.
Result
The initial step of the assay verification and validation pathway was to measure the performance of each assay in a standard curve titration analysis. Key metrics are curve fit (R-square values) showing quality of technical replicates and amplification dynamic as well as slope of the regression line to calculate PCR efficiencies. All four novel assays for the detection of DIV1 (DIV-CSG, -PAT, -PAP, and -ATPase) show a tight curve fit with R-Square values greater0.99 along a dynamic range of 1 to 10,000 copies per reaction (Figure 1). The amplification efficiency for each of the assays is, DIV PAP 104.3%, DIV ATPase 96.3%, DIV CSG 95.6%, and DIV PAT 93.8% (Table 3). All four assays werefurther validated with respect to their analytical performance. For analytical specificity (ASp) samples positive for DIV1 were confirmed positive (100% inclusivity) and direct amplicon sequencing and database interrogation employed to validate the confirmed results (Table 4). Moreover, L. vannamei and P. monodon samples negative for DIV1 but positive for other shrimp pathogens such as HPV, IHHNV, MBV, LSNV, Vibrio parahaemolyticus (PirA/PirB) and EHP showed no false positive DIV1 assay results (exclusivity). No cross-reactivity of all new qPCR assays has also been confirmed with polychaete worm samples from the genus Perinereis.Analytical sensitivity or LOD was determined to be 2.2 copies/reaction (Ct = 32.0; DIV PAT), 2.9 copies/ reaction (Ct = 32.1; DIV CSG), 3.1 copies/reaction (Ct = 31.3; DIV PAP), and 8.6 copies/reaction (Ct = 32.2; DIV ATPase) (Table 5).
Diagnostic metrics sensitivity (DSe) and DSp were based on the traditional golden reference assay approach with each assay evaluated as golden reference point to determine the differential DSe and DSp values for each of the novel DIV1 assays using a single population of 91 L. vannamei shrimp samples known to be infected with DIV1. The binary qPCR profile for all samples (Table 6) was established and formed the basis for the DSe/DSp assessment. With DIV ATPase set as golden reference assay 100% DSe/100% DSp, the results for the other assay were obtained as 97%/93.1% (DIV CSG), 93.9%/93.1% (DIV PAP), and 100%/93.1% (DIV PAT), respectively. Setting DIV CSG as reference 100% DSe/100% DSp, all comparative assays were 88.9%/98.2% (DIV ATPase), 91.7%/96.4% (DIV PAP), and 97.2%/96.4% (DIV PAT). DIV PAP as reference yielded DSe/DSp results 88.6%/96.4% for DIV ATPase, 94.3%/94.6% for DIV CSG, and 97.1%/94.6% for DIV PAT. Finally, DIV PAT as golden reference assay resulted in a DSe/DSp of 89.2%/100% for DIV ATPase, 94.6%/98.1%, for DIV CSG, and 91.9%/98.1% for DIV PAT.
Conclusion
DIV1 is a critical emerging pathogen threat for the global shrimp industry, and all necessary steps need to be taken to avoid spread of the disease into the Asia-Pacific region and beyond. Hence, one key approach is to develop novel target gene and assays for this emerging pathogen and increase molecular tool capability for national and international biosecurity efforts. The novel targets and assays presented in this study are a vital step toward the tool expansion and will serve as a key foundation for increasing biosecurity preparedness. In particular, the DIV PAT and DIV PAP assays provide two new tools with high specificity and sensitivity toward monitoring and detecting this pathogen.
Melony J. Sellars, Louise Franz, Ralf Joachim Moser
Genics Pty Ltd, St Lucia, Queensland, Australia






